Validation - pesticide use values
Paige Baisley + Maggie Douglas
Purpose
This code compares pesticide use estimates across independent datasets for major crops (corn, soy, wheat, cotton, rice). The comparisons include:
- total amount applied (kg)
- application rate (kg/ha) calculated by dividing the USDA total kg by our new estimate of crop area (ha)
- application rate (kg/ha) derived indirectly by multiplying the USDA average application rate on treated hectares by the percent of area treated
Libraries & functions
library(tidyverse)
library(readxl)
library(dplyr)
library(ggplot2)
library(gridExtra)Load and join data
## Key relating USDA and USGS pesticide names
pesticide_names <- read_excel("../keys/usda_usgs_pesticide_names.xlsx")
# check for duplicates
usgs_pesticide_dups <- pesticide_names %>%
group_by(usgs_cmpd) %>%
mutate(n = n()) %>%
filter(Match_status != "missing")
# some USGS names correspond to > 1 USDA name
usda_pesticide_dups <- pesticide_names %>%
group_by(usda_cmpd) %>%
mutate(n = n())
# no true duplicates in USDA names (some extra spaces, etc.)
## USGS pesticide estimates
usgs <- read.csv("../output_big/bee_tox_index_state_yr_cmpd_20200609.csv") %>%
mutate(crop = toupper(crop))
## USDA pesticide estimates
usda <- read.csv("../output_big/nass_survey/qs.environment.pest.st.app.rate.usgs_20200528.csv")
# Join USDA estimates and name key
usda_key <- left_join(usda, pesticide_names, by = c("DOMAINCAT_DESC" = "usda_cmpd"))%>%
mutate(kg_usda = lb*0.453592)
## Sum kg_usda values for data with repeated USGS cmpd, year, crop, and state
# aggregate cases in which multiple USDA AIs correspond to one USGS AI
usda_agg <- usda_key %>%
group_by(YEAR, STATE_ALPHA, usgs_cmpd, COMMODITY_DESC, CLASS_DESC, usda_category) %>%
summarise(n_usda = length(kg_usda),
kg_usda = sum(kg_usda, na.rm = TRUE),
kg_ha_yr_ave = sum(kg_ha_yr_ave, na.rm = TRUE),
perc_trt_ave = sum(perc_trt_ave, na.rm = TRUE)) %>%
mutate(perc_trt_alt = ifelse(perc_trt_ave <= 100, perc_trt_ave, 100)) %>% # create new % trt column bounded at 100%
rename(CROP = COMMODITY_DESC) %>%
ungroup()## `summarise()` has grouped output by 'YEAR', 'STATE_ALPHA', 'usgs_cmpd', 'COMMODITY_DESC', 'CLASS_DESC'. You can override using the `.groups` argument.usda_agg <- usda_agg[!is.na(usda_agg$usgs_cmpd), ] # remove rows without USGS compound
# aggregate cases in which multiple values for a given crop-state-year-AI combo (only occurs in wheat)
# this is imperfect b/c ideally the averages would be weighted by acreage
usda_agg_other <- filter(usda_agg, CROP != "WHEAT") %>%
select(-CLASS_DESC) %>%
mutate(n_crop = 1) %>%
select(YEAR, STATE_ALPHA, usgs_cmpd, CROP, usda_category, n_usda, n_crop, kg_usda, kg_ha_yr_ave, perc_trt_ave, perc_trt_alt)
usda_agg_wheat <- filter(usda_agg, CROP == "WHEAT") %>%
group_by(YEAR, STATE_ALPHA, usgs_cmpd, CROP, usda_category) %>%
summarise(n_usda = sum(n_usda),
n_crop = length(kg_usda),
kg_usda = sum(kg_usda, na.rm = TRUE),
kg_ha_yr_ave = mean(kg_ha_yr_ave, na.rm = TRUE),
perc_trt_ave = mean(perc_trt_ave, na.rm = TRUE),
perc_trt_alt = mean(perc_trt_alt, na.rm = TRUE)) %>%
ungroup()## `summarise()` has grouped output by 'YEAR', 'STATE_ALPHA', 'usgs_cmpd', 'CROP'. You can override using the `.groups` argument.usda_full <- bind_rows(usda_agg_other, usda_agg_wheat)
## Join aggregated USDA estimates and USGS estimate
joined <- inner_join(usda_full, usgs,
by=c("usgs_cmpd"="cmpd_usgs",
"STATE_ALPHA"="STATE_ALPHA",
"YEAR"="Year",
"CROP"="crop"))%>%
mutate(kg_ha_usda = kg_usda/ha,
kg_ha2_usda = (kg_ha_yr_ave),
kg_thou_usda = kg_usda/10^3,
kg_thou_usgs = kg_app/10^3)
joined[joined == 0] <- NA # restore NA valuesSubset and clean data
# Select for desired years and crops
crops <- c("CORN", "SOYBEANS", "WHEAT", "COTTON", "RICE")
target <- c(1997:2017)
category <- c("Insecticide", "Herbicide", "Fungicide")
# Prep dataset
clean <- joined %>%
filter(CROP %in% crops,
YEAR %in% target,
usda_category %in% category)%>%
mutate(kg_ha_usgs = kg_ha)%>%
select(kg_app,
kg_usda,
kg_thou_usda,
kg_thou_usgs,
kg_ha_usda,
kg_ha2_usda,
kg_ha_usgs,
CROP,
usda_category,
STATE_ALPHA,
YEAR,
usgs_cmpd,
n_usda,
n_crop,
perc_trt_alt,
ha)
# create groups for percent treated
perc_grp_tags <- c("10","20", "30", "40", "50", "60", "70", "80","90", "90+")
diff <- clean %>%
mutate(kg_diff = kg_app - kg_usda,
kg_ha_diff1 = kg_ha_usgs - kg_ha_usda,
kg_ha_diff2 = kg_ha_usgs - kg_ha2_usda,
RE_kg = (kg_diff/((kg_app + kg_usda)/2))*100,
RE_kg_ha_diff1 = (kg_ha_diff1/((kg_ha_usgs + kg_ha_usda)/2))*100,
RE_kg_ha_diff2 = (kg_ha_diff2/((kg_ha_usgs + kg_ha2_usda)/2))*100) %>%
mutate(perc_grp = case_when(
perc_trt_alt < 10 ~ perc_grp_tags[1],
perc_trt_alt >= 10 & perc_trt_alt < 20 ~ perc_grp_tags[2],
perc_trt_alt >= 20 & perc_trt_alt < 30 ~ perc_grp_tags[3],
perc_trt_alt >= 30 & perc_trt_alt < 40 ~ perc_grp_tags[4],
perc_trt_alt >= 40 & perc_trt_alt < 50 ~ perc_grp_tags[5],
perc_trt_alt >= 50 & perc_trt_alt < 60 ~ perc_grp_tags[6],
perc_trt_alt >= 60 & perc_trt_alt < 70 ~ perc_grp_tags[7],
perc_trt_alt >= 70 & perc_trt_alt < 80 ~ perc_grp_tags[8],
perc_trt_alt >= 80 & perc_trt_alt < 90 ~ perc_grp_tags[9],
perc_trt_alt >= 90 ~ perc_grp_tags[10]
))Visualize patterns
Difference by area treated
ggplot(data = drop_na(diff, perc_trt_alt), mapping = aes(x = perc_grp, y = RE_kg)) +
geom_jitter(color = "darkturquoise", alpha=0.1) +
geom_boxplot(fill="ivory3", color="black", alpha=0.3) +
geom_hline(yintercept = 0, lty = 2) +
ggtitle('a) kg applied') +
labs(x = '% crop area treated (USDA)',
y = 'Relative difference (%)') +
guides(color=FALSE) +
theme_minimal(base_size = 18)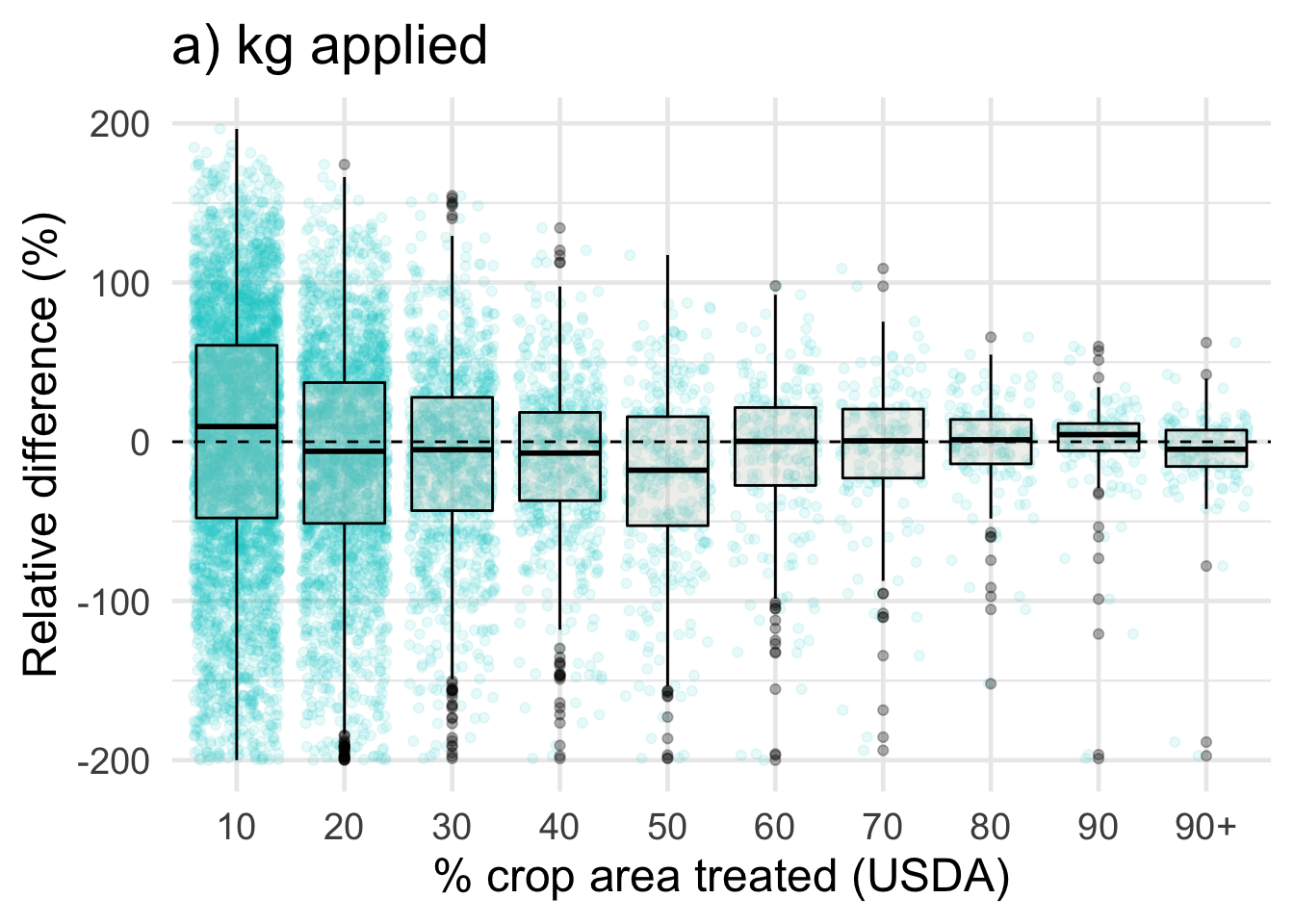
ggplot(data = drop_na(diff, perc_trt_alt), mapping = aes(x = perc_grp, y = RE_kg_ha_diff1)) +
geom_jitter(color = "mediumorchid1", alpha=0.2) +
geom_boxplot(fill="ivory3", color="black", alpha=0.2) +
geom_hline(yintercept = 0, lty = 2) +
ggtitle('b) kg/ha - Method 1') +
labs(x = '% crop area treated (USDA)',
y = 'Relative difference (%)') +
guides(color=FALSE) +
theme_minimal(base_size = 18)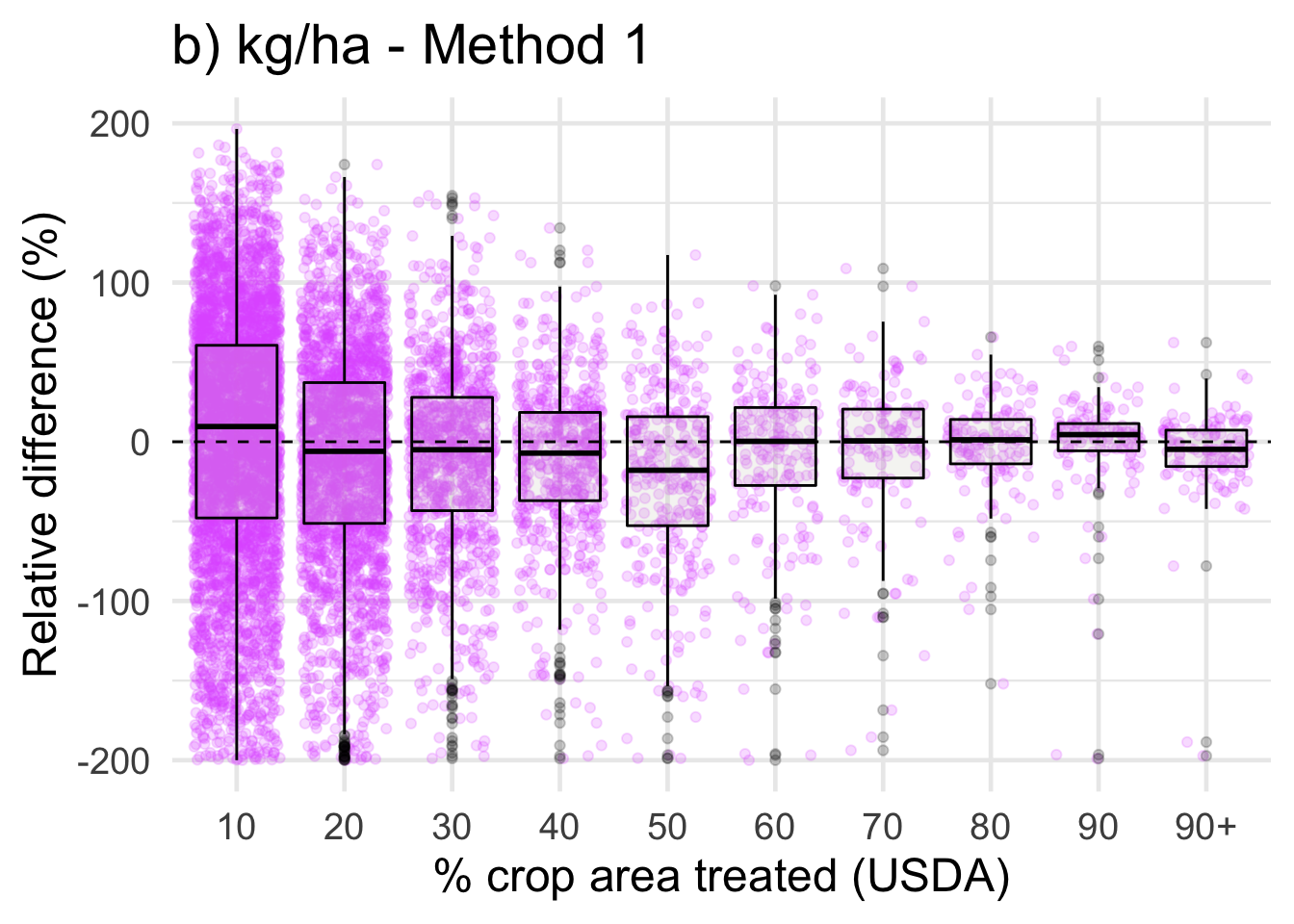
ggplot(data = drop_na(diff, perc_trt_alt), mapping = aes(x = perc_grp, y = RE_kg_ha_diff2)) +
geom_jitter(color = "coral1", alpha=0.1) +
geom_boxplot(fill="ivory3", color="black", alpha=0.3) +
geom_hline(yintercept = 0, lty = 2) +
ggtitle('c) kg/ha - Method 2') +
labs(x = '% crop area treated (USDA)',
y = 'Relative difference (%)') +
guides(color=FALSE) +
theme_minimal(base_size = 18)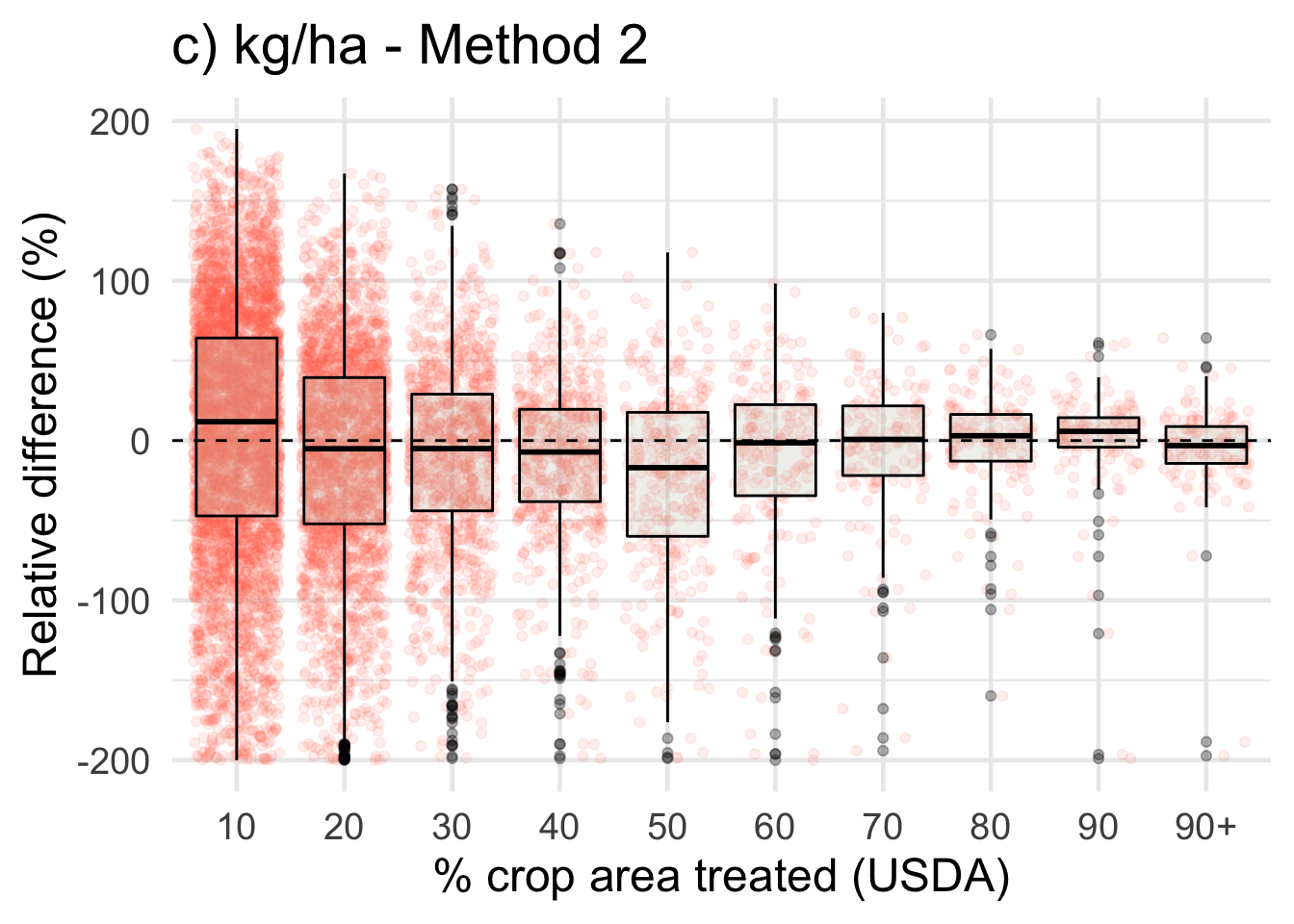
Save figure for publication
require(gridExtra)
a <- ggplot(data = drop_na(diff, perc_trt_alt), mapping = aes(x = perc_grp, y = RE_kg)) +
geom_jitter(color = "darkturquoise", alpha=0.1) +
geom_boxplot(fill="ivory3", color="black", alpha=0.3) +
geom_hline(yintercept = 0, lty = 2) +
ggtitle('a) kg applied') +
labs(x = '% crop area treated (USDA)',
y = 'Relative difference (%)') +
guides(color="none") +
theme_minimal(base_size = 18)
b <- ggplot(data = drop_na(diff, perc_trt_alt), mapping = aes(x = perc_grp, y = RE_kg_ha_diff1)) +
geom_jitter(color = "mediumorchid1", alpha=0.2) +
geom_boxplot(fill="ivory3", color="black", alpha=0.2) +
geom_hline(yintercept = 0, lty = 2) +
ggtitle('b) kg/ha - Method 1') +
labs(x = '% crop area treated (USDA)',
y = 'Relative difference (%)') +
guides(color="none") +
theme_minimal(base_size = 18)
c <- ggplot(data = drop_na(diff, perc_trt_alt), mapping = aes(x = perc_grp, y = RE_kg_ha_diff2)) +
geom_jitter(color = "coral1", alpha=0.1) +
geom_boxplot(fill="ivory3", color="black", alpha=0.3) +
geom_hline(yintercept = 0, lty = 2) +
ggtitle('c) kg/ha - Method 2') +
labs(x = '% crop area treated (USDA)',
y = 'Relative difference (%)') +
guides(color="none") +
theme_minimal(base_size = 18)
ggsave("../docs/validation_values_files/figure-html/fig3.pdf",
arrangeGrob(a,b,c),
width = 3,
height = 9,
scale = 1.8)Difference by area treated and crop
ggplot(data = drop_na(diff, perc_trt_alt), mapping = aes(x = perc_grp, y = RE_kg)) +
geom_jitter(aes(color = CROP), alpha=0.2) +
geom_boxplot(fill="bisque", color="black", alpha=0.3) +
geom_hline(yintercept = 0, lty = 2) +
ggtitle('kg applied') +
labs(x = '% crop area treated (USDA)',
y = 'Relative difference (%)') +
facet_grid(~ CROP) +
guides(color=FALSE) +
theme_minimal()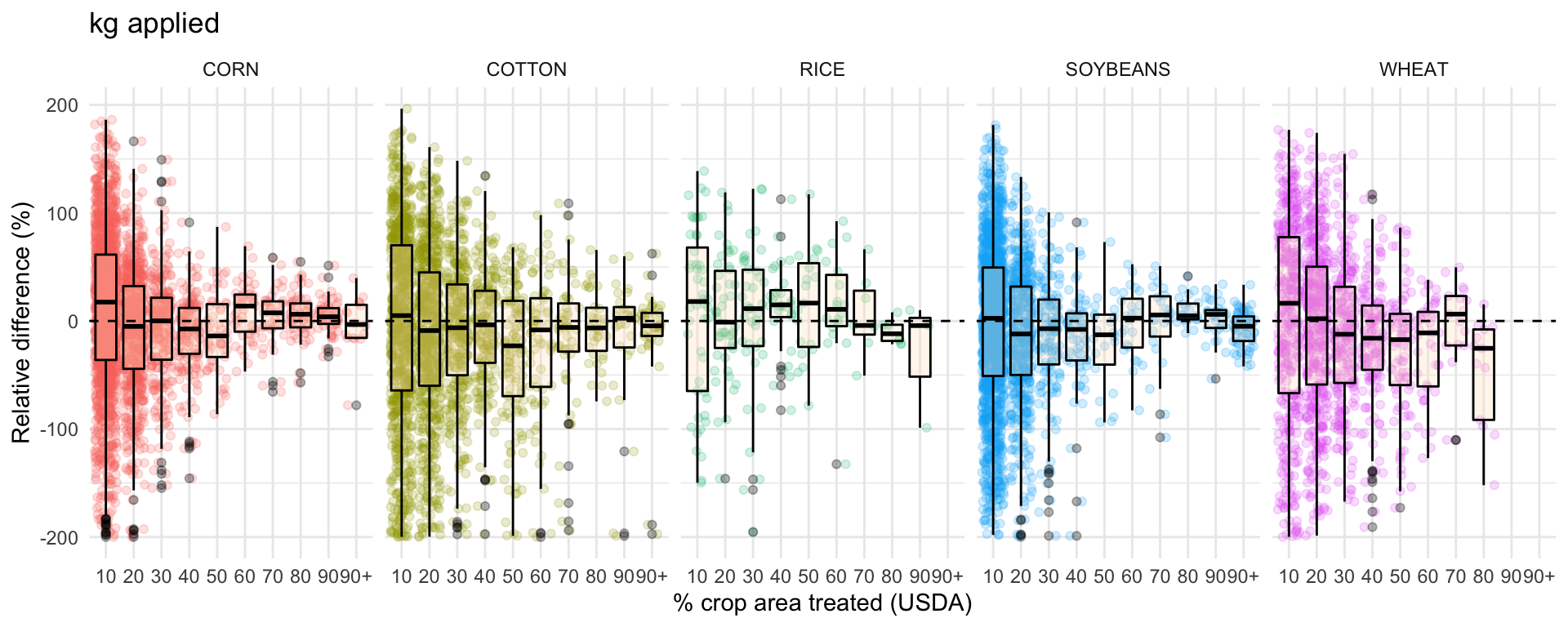
ggplot(data = drop_na(diff, perc_trt_alt), mapping = aes(x = perc_grp, y = RE_kg_ha_diff1)) +
geom_jitter(aes(color = CROP), alpha=0.2) +
geom_boxplot(fill="bisque", color="black", alpha=0.3) +
geom_hline(yintercept = 0, lty = 2) +
ggtitle('kg/ha - Method 1') +
labs(x = '% crop area treated (USDA)',
y = 'Relative difference (%)') +
facet_grid(~ CROP) +
guides(color=FALSE) +
theme_minimal()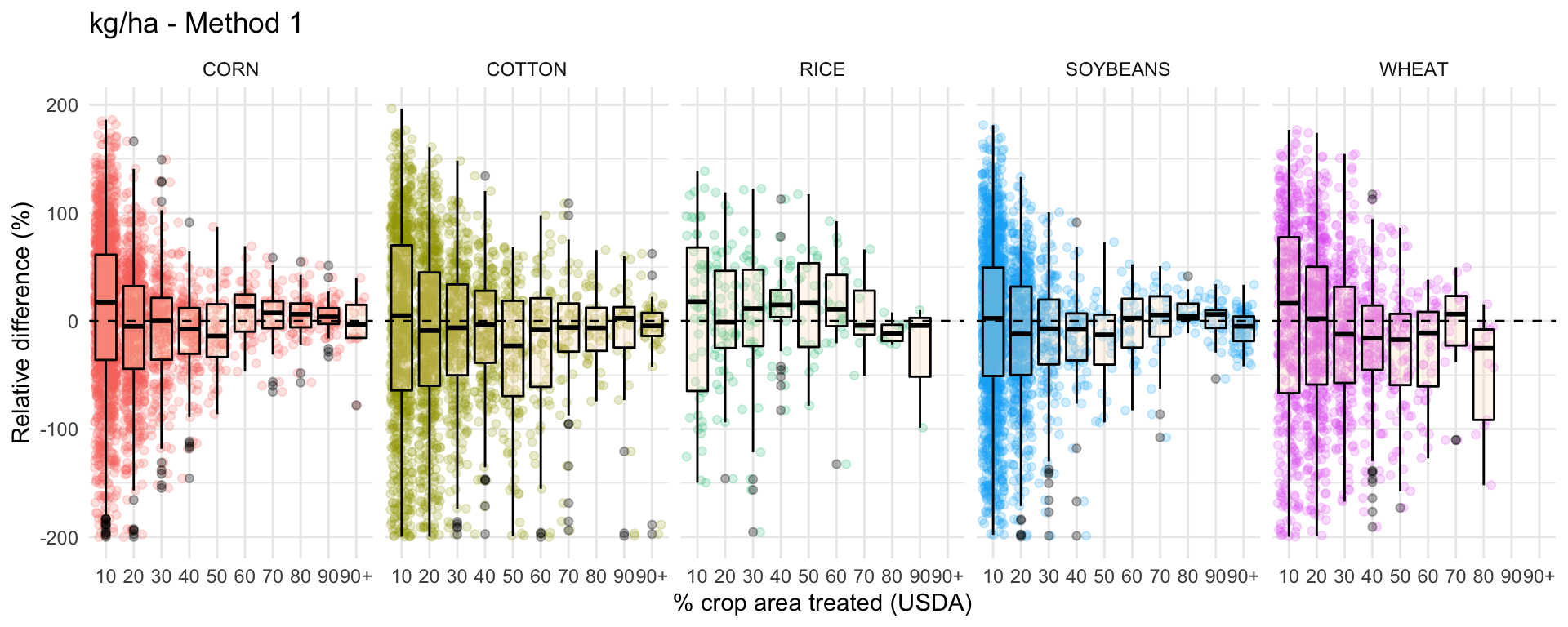
ggplot(data = drop_na(diff, perc_trt_alt), mapping = aes(x = perc_grp, y = RE_kg_ha_diff2)) +
geom_jitter(aes(color = CROP), alpha=0.2) +
geom_boxplot(fill="bisque", color="black", alpha=0.3) +
geom_hline(yintercept = 0, lty = 2) +
ggtitle('kg/ha - Method 2') +
labs(x = '% crop area treated (USDA)',
y = 'Relative difference (%)') +
facet_grid(~ CROP) +
guides(color=FALSE) +
theme_minimal()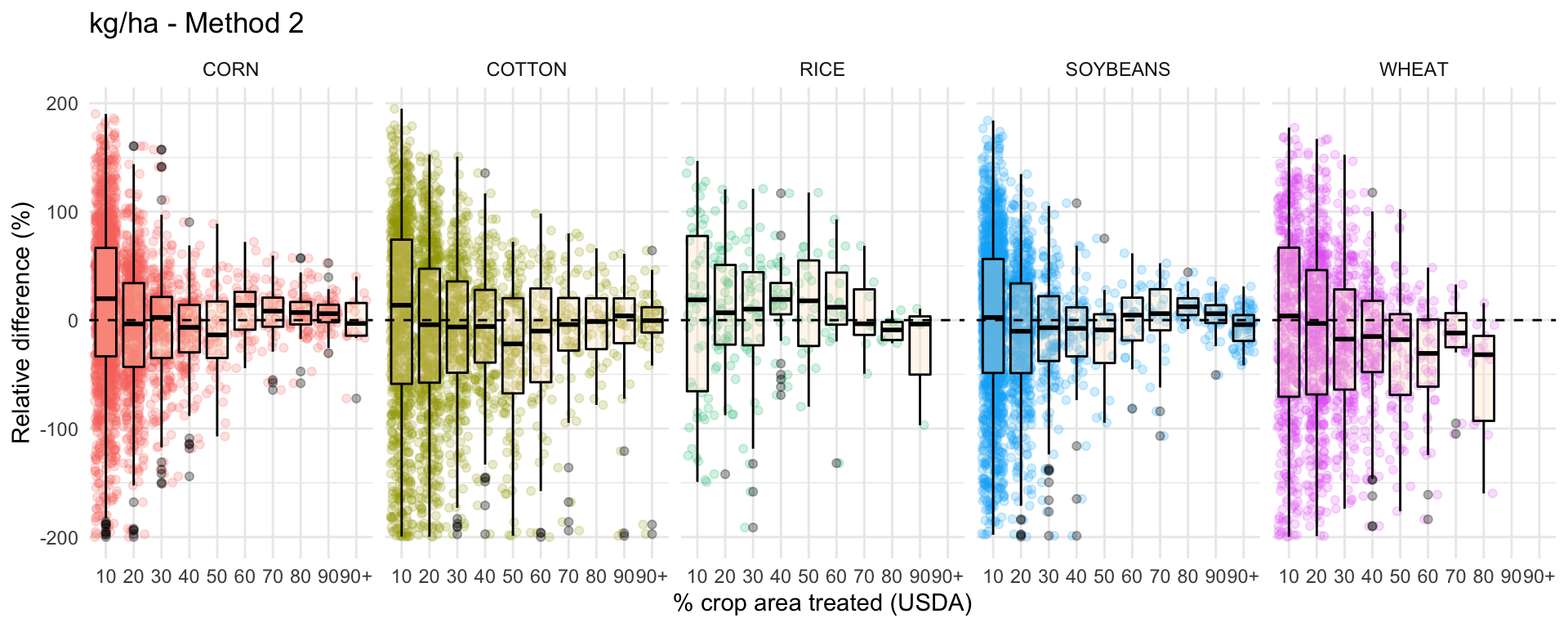
Restructure for additional plotting
# Restructure corrected estimates data for plots
# rename USGS data columns according to the method of analysis their values will be used in
names(clean)[names(clean)=="kg_thou_usgs"] <- "kg_thou"
names(clean)[names(clean)=="kg_ha_usgs"] <- "method_1"
# create data set with only USGS values
# duplicate and rename method 1 values so that they can be used for method 2 analysis
# turn to long form
usgs_cl <- clean %>%
select(kg_thou,
method_1,
CROP,
STATE_ALPHA,
YEAR,
usgs_cmpd,
usda_category)%>%
mutate(method_2 = method_1) %>%
gather("kg_thou","method_1","method_2",
key = "method",
value = "usgs")
# create data set with only USDA values
# turn to long form
usda_cl <- clean %>%
select(kg_thou_usda,
kg_ha_usda,
kg_ha2_usda,
STATE_ALPHA,
YEAR,
usgs_cmpd,
CROP,
usda_category) %>%
rename(kg_thou = kg_thou_usda,
method_1 = kg_ha_usda,
method_2 = kg_ha2_usda) %>%
gather("kg_thou", "method_1","method_2",
key = "method",
value = "usda")
# join altered usgs and usda data with single methods column;
# unite methods and usda category to allow facet wrap to be used
graph_data <- full_join(usda_cl, usgs_cl,
by=c("CROP",
"STATE_ALPHA",
"YEAR",
"method",
"usgs_cmpd",
"usda_category"))%>%
unite("methods", c("method", "usda_category"), remove = FALSE) %>%
mutate(diff = usgs-usda,
RE = (diff/((usda+usgs)/2)*100))
graph_data$method_pretty <- recode(graph_data$methods,
kg_thou_Fungicide = "Fungicides (1000 kg)",
kg_thou_Insecticide = "Insecticides (1000 kg)",
kg_thou_Herbicide = "Herbicides (1000 kg)",
method_1_Fungicide = "Fungicides (kg/ha - Method 1)",
method_1_Herbicide = "Herbicides (kg/ha - Method 1)",
method_1_Insecticide = "Insecticides (kg/ha - Method 1)",
method_2_Fungicide = "Fungicides (kg/ha - Method 2)",
method_2_Herbicide = "Herbicides (kg/ha - Method 2)",
method_2_Insecticide = "Insecticides (kg/ha - Method 2)")
# export comparison dataset for low values
write.csv(graph_data, "../output_big/validation_low_estimate.csv", row.names = FALSE)Plot estimates against each other
# plot by methods of analysis and USDA category;
# create invisible matching points so that each graph x-axis=y-axis
ggplot(graph_data, aes(x=usda, y=usgs, color=CROP))+
geom_point(mapping=aes(x=usda,
y=usgs),
alpha = 0.4)+
geom_point(mapping=aes(x=usgs,
y=usda),
alpha=0)+
geom_abline(linetype = 2)+
facet_wrap("methods", scales="free") +
theme_classic() +
xlab("USDA estimate") +
ylab("USGS-based estimate")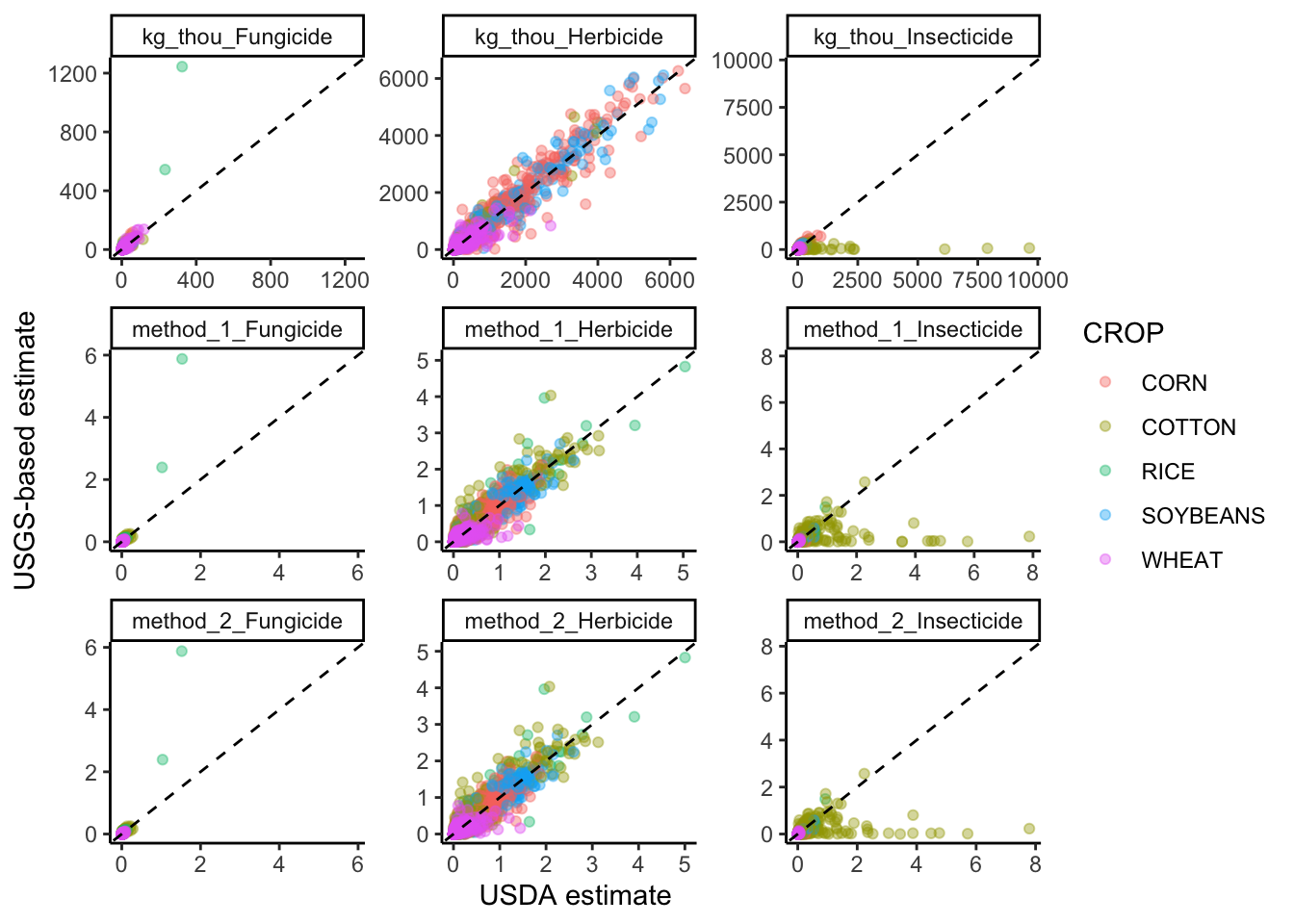
# remove outliers
ggplot(filter(graph_data,
!(CROP == "COTTON" & usgs_cmpd == "MALATHION")
& !(CROP == "RICE" & usgs_cmpd == "COPPERSULFATE")),
aes(x=usda, y=usgs, color=CROP))+
geom_point(mapping=aes(x=usda,
y=usgs),
alpha = 0.4)+
geom_point(mapping=aes(x=usgs,
y=usda),
alpha=0)+
geom_abline(linetype = 2)+
facet_wrap("method_pretty", scales="free") +
xlab("USDA estimate") +
ylab("USGS-based estimate") +
theme_classic() +
theme(strip.background = element_blank()) 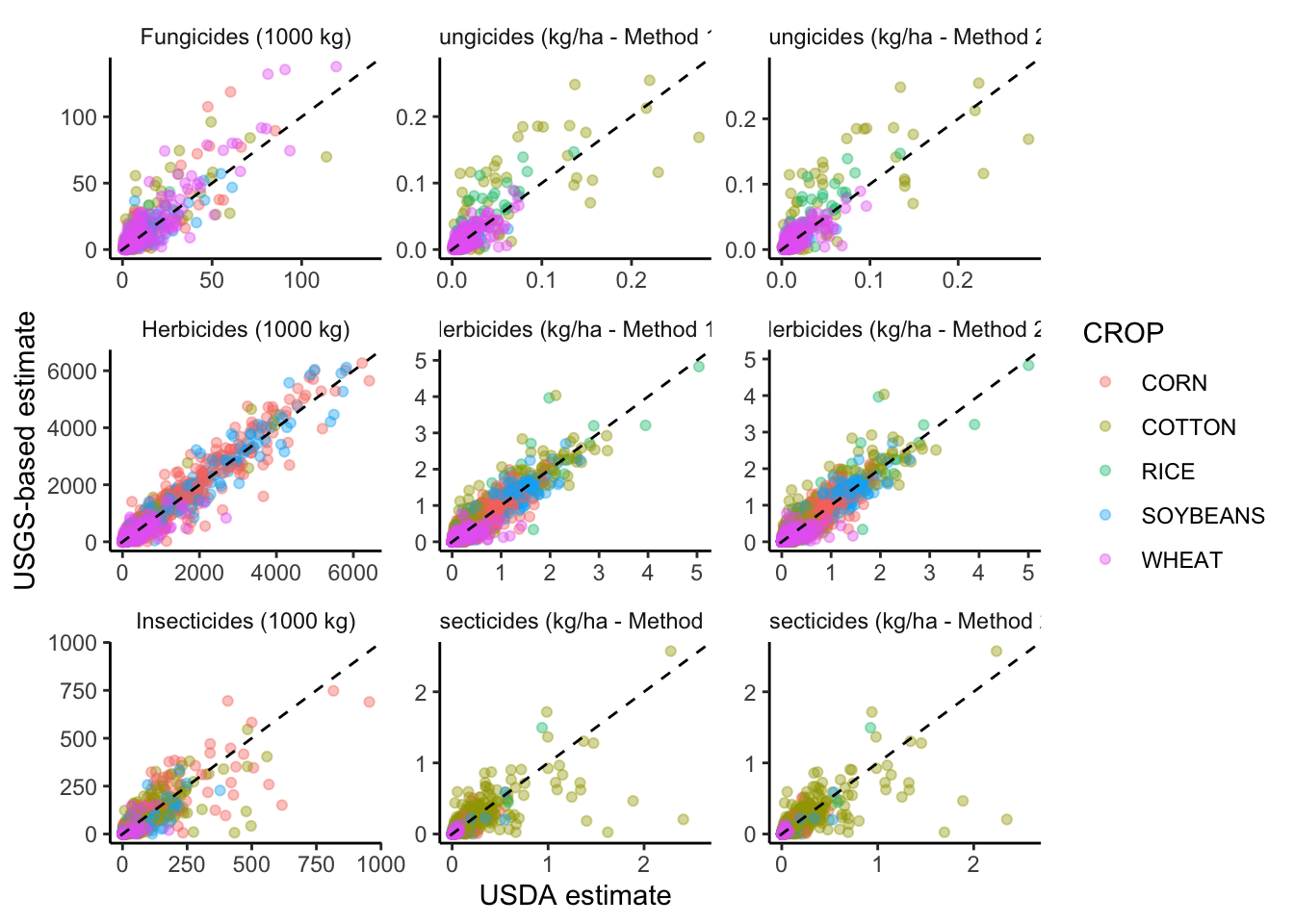
Save figure for publication
# open file
pdf(file = "../docs/validation_values_files/figure-html/fig2.pdf", # The directory you want to save the file in
width = 8, # The width of the plot in inches
height = 6) # The height of the plot in inches
# remove outliers
ggplot(filter(graph_data,
!(CROP == "COTTON" & usgs_cmpd == "MALATHION")
& !(CROP == "RICE" & usgs_cmpd == "COPPERSULFATE")),
aes(x=usda, y=usgs, color=CROP))+
geom_point(mapping=aes(x=usda,
y=usgs),
alpha = 0.4)+
geom_point(mapping=aes(x=usgs,
y=usda),
alpha=0)+
geom_abline(linetype = 2)+
facet_wrap("method_pretty", scales="free") +
xlab("USDA estimate") +
ylab("USGS-based estimate") +
theme_classic() +
theme(strip.background = element_blank())
# Run dev.off() to create the file!
dev.off()## quartz_off_screen
## 2Boxplot of RE by class
# plot by methods of analysis and USDA category;
# create invisible matching points so that each graph x-axis=y-axis
ggplot(graph_data, aes(x=method, y=RE, color=usda_category))+
geom_boxplot() +
geom_hline(yintercept = 0, linetype = 2) +
#facet_wrap("method", scales="free") +
theme_classic() +
xlab("") +
ylab("Relative difference (%)")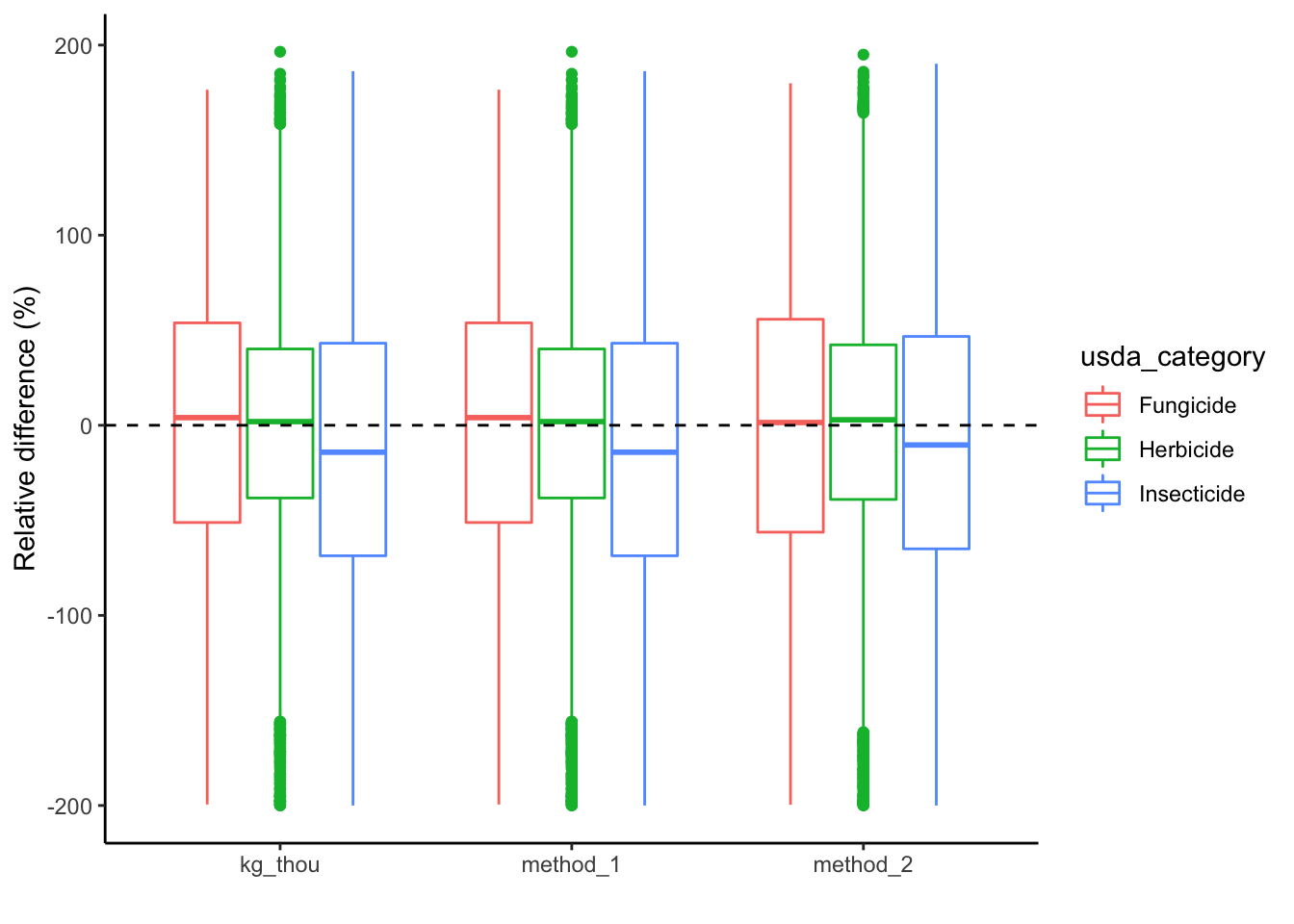
Boxplot of RE by crop + class
# plot by methods of analysis and USDA category;
# create invisible matching points so that each graph x-axis=y-axis
ggplot(graph_data, aes(x=CROP, y=RE, color=CROP))+
geom_boxplot() +
geom_hline(yintercept = 0, linetype = 2) +
facet_wrap("methods", scales="free") +
theme_classic() +
xlab("") +
ylab("Relative difference (%)")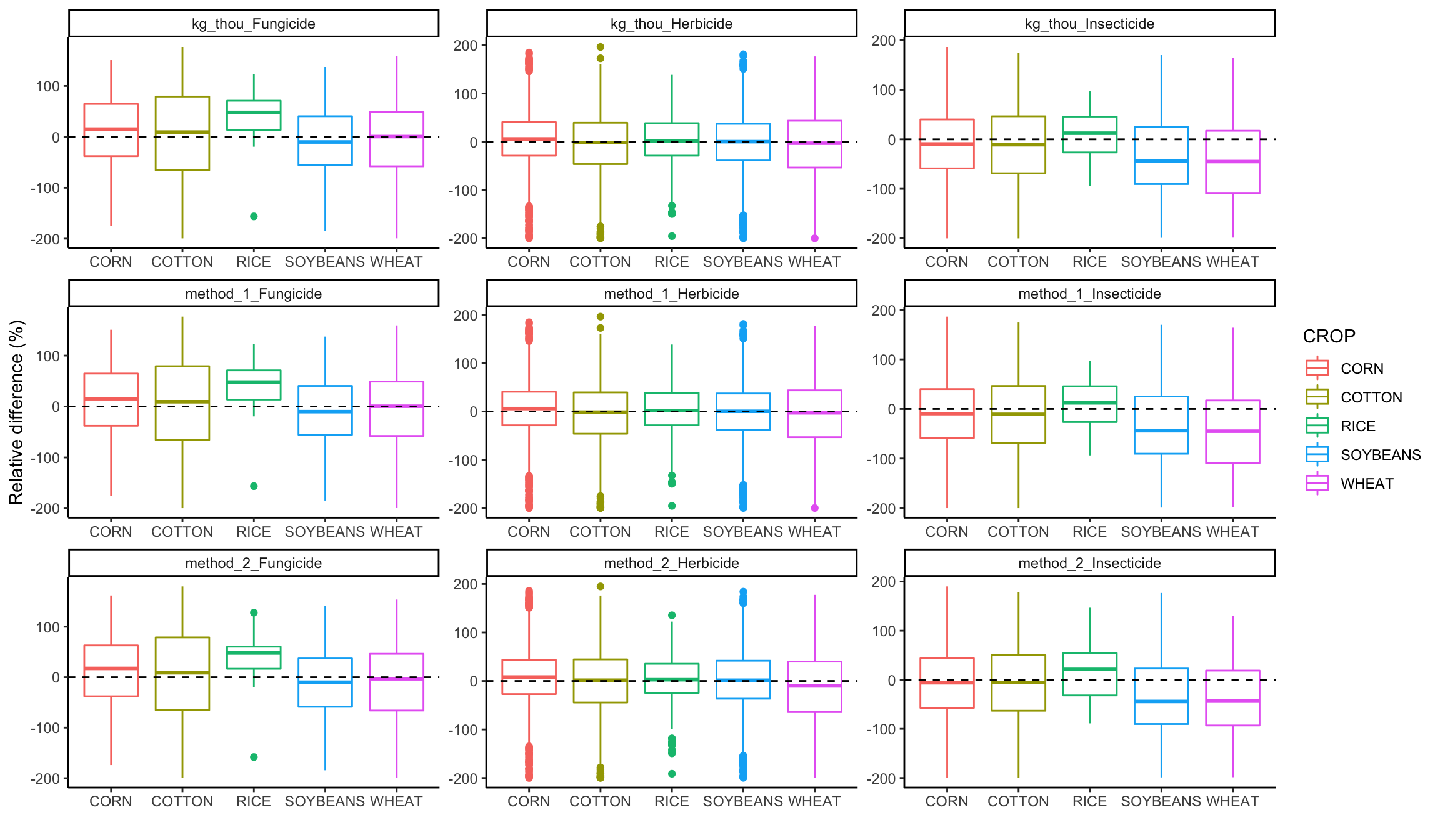
Boxplot of RE by year
# plot by methods of analysis and USDA category;
# create invisible matching points so that each graph x-axis=y-axis
ggplot(graph_data, aes(x=as.factor(YEAR), y=RE))+
geom_boxplot() +
geom_hline(yintercept = 0, linetype = 2) +
facet_wrap("CROP", scales="free") +
theme_classic() +
xlab("") +
ylab("Relative difference (%)")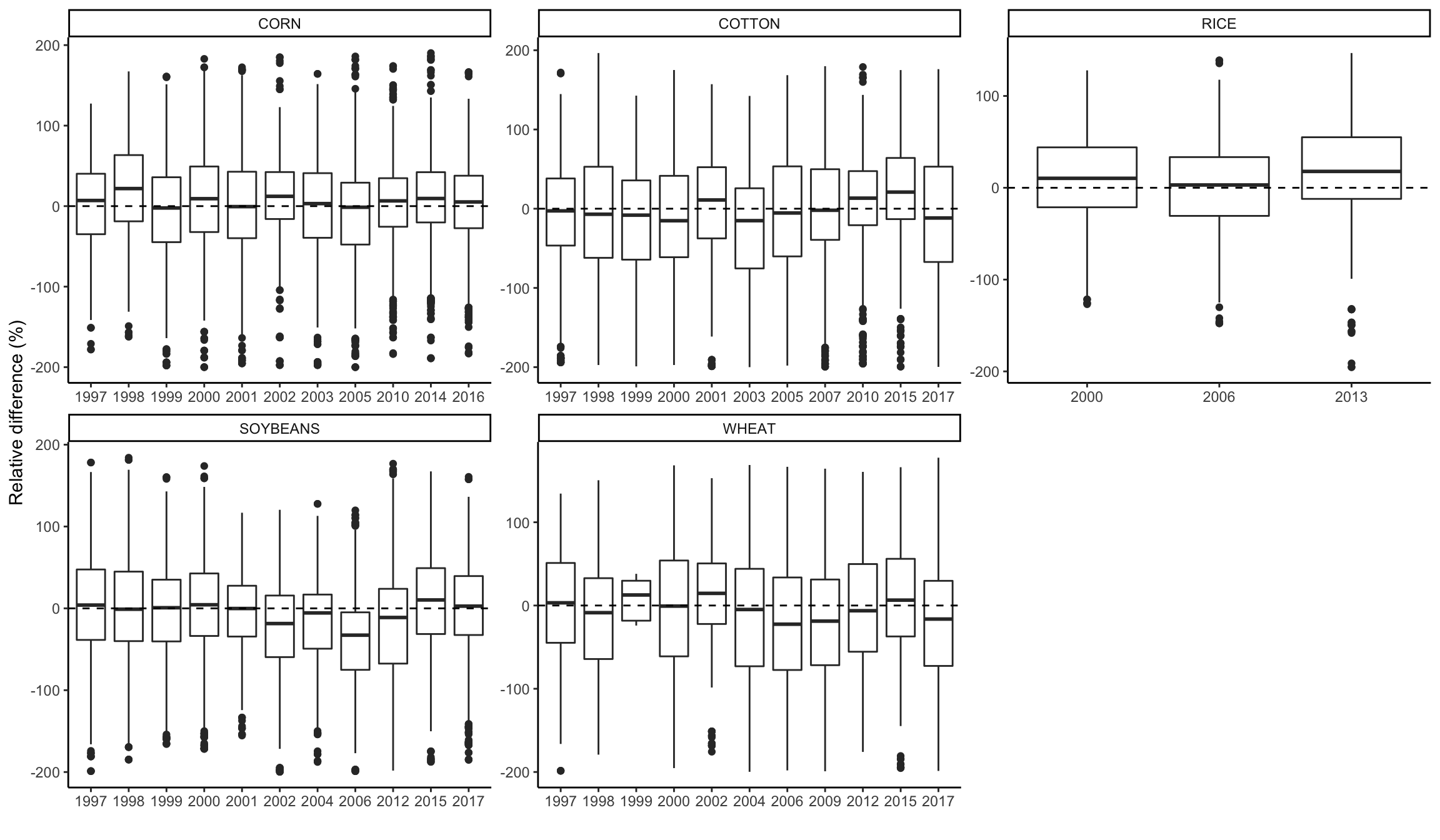
Example trends comparison
# e.g. for glyphosate
# plot by methods of analysis and USDA category;
# create invisible matching points so that each graph x-axis=y-axis
filtered_data <- graph_data %>%
filter(methods == "method_2_Herbicide",
usgs_cmpd == "GLYPHOSATE",
CROP == "SOYBEANS",
!STATE_ALPHA %in% c("MD","DE")) %>%
select(-diff, -RE) %>%
pivot_longer(cols = usda:usgs,
names_to = "source",
values_to = "kg_ha")
ggplot(filtered_data, aes(x = YEAR, y = kg_ha, group = source)) +
geom_line(aes(linetype = source)) +
facet_wrap("STATE_ALPHA", scales = "free_y") +
theme_classic() +
xlab("") +
ylab("Application rate (kg/ha)")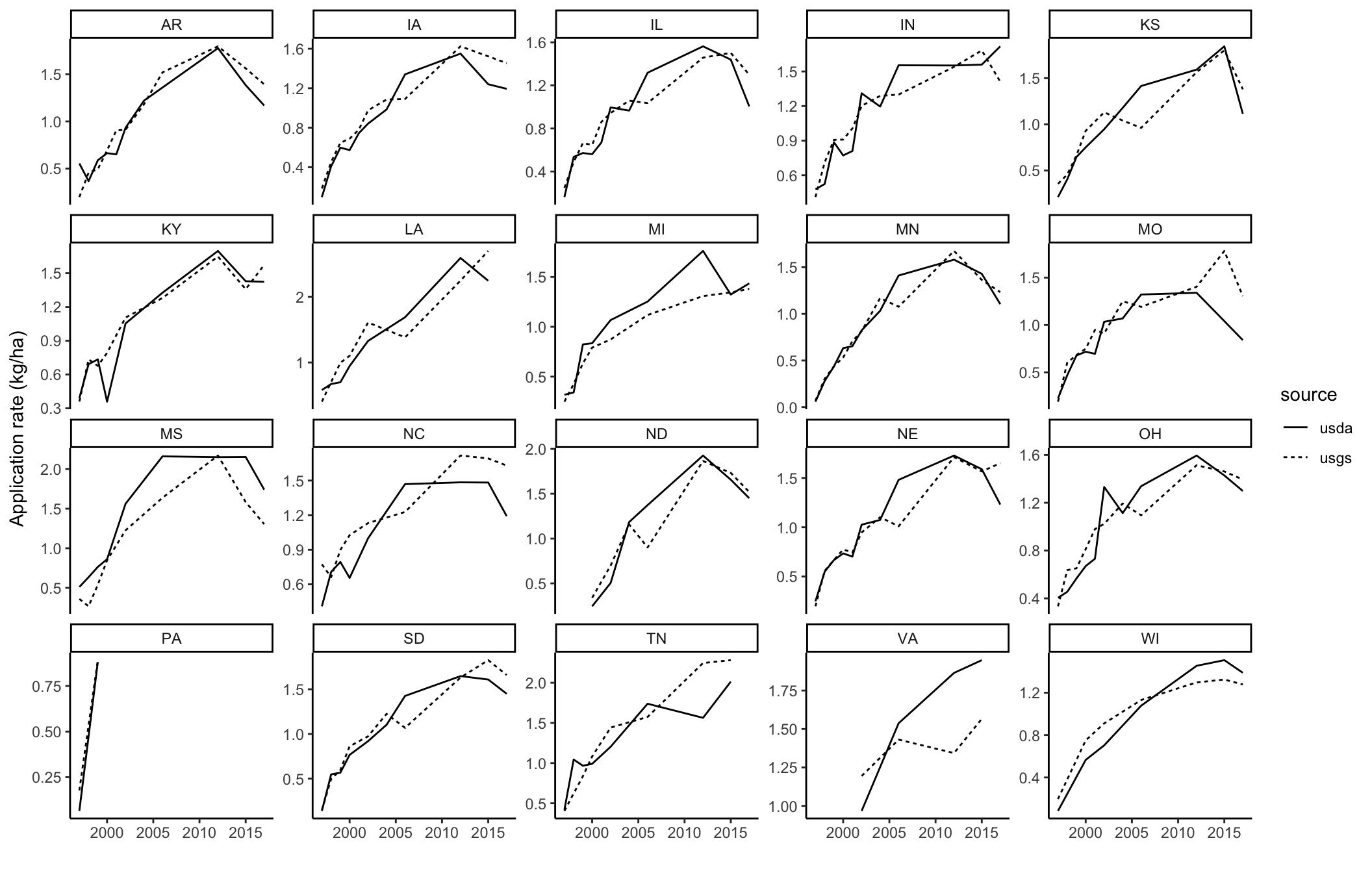
Calculate summary statistics
By category
stats <- graph_data %>%
drop_na() %>%
group_by(usda_category, method) %>%
summarise(n = n(),
med_RE = round(median(RE, na.rm=T), digits = 1),
iqr_25 = round(quantile(RE, .25, na.rm=T), digits = 1),
iqr_75 = round(quantile(RE, .75, na.rm=T), digits = 1),
wilcox_p = (wilcox.test(RE, mu = 0, alternative = "two.sided", conf.int=T, conf.level = 0.05))$p.value,
rank_corr = round(cor(usgs, usda, method = "spearman", use = "pairwise.complete.obs"), digits = 2),
corr = round(cor(usgs, usda, method = "pearson", use = "pairwise.complete.obs"), digits = 2)
)## `summarise()` has grouped output by 'usda_category'. You can override using the `.groups` argument.`%notin%` <- Negate(`%in%`)
out_removed <- filter(graph_data,
!(CROP == "COTTON" & usgs_cmpd == "MALATHION"))
stats_out_removed <- out_removed %>%
drop_na() %>%
group_by(usda_category, method) %>%
summarise(n = n(),
med_RE = round(median(RE, na.rm=T), digits = 1),
iqr_25 = round(quantile(RE, .25, na.rm=T), digits = 1),
iqr_75 = round(quantile(RE, .75, na.rm=T), digits = 1),
wilcox = (wilcox.test(RE, mu = 0, alternative = "two.sided"))$p.value,
rank_corr = round(cor(usgs, usda, method = "spearman", use = "pairwise.complete.obs"), digits = 2),
corr = round(cor(usgs, usda, method = "pearson", use = "pairwise.complete.obs"), digits = 2))## `summarise()` has grouped output by 'usda_category'. You can override using the `.groups` argument.stats_total <- graph_data %>%
group_by(method) %>%
drop_na() %>%
summarise(n = n(),
med_RE = round(median(RE, na.rm=T), digits = 1),
iqr_25 = round(quantile(RE, .25, na.rm=T), digits = 1),
iqr_75 = round(quantile(RE, .75, na.rm=T), digits = 1),
wilcox = (wilcox.test(RE, mu = 0, alternative = "two.sided"))$p.value,
rank_corr = round(cor(usgs, usda, method = "spearman", use = "pairwise.complete.obs"), digits = 2),
corr = round(cor(usgs, usda, method = "pearson", use = "pairwise.complete.obs"), digits = 2))By compound
stats_cmpd <- graph_data %>%
drop_na() %>%
group_by(usgs_cmpd, usda_category, method) %>%
summarise(med_RE = round(median(RE, na.rm=T), digits = 1),
iqr_25 = round(quantile(RE, .25, na.rm=T), digits = 1),
iqr_75 = round(quantile(RE, .75, na.rm=T), digits = 1),
rank_corr = round(cor(usgs, usda, method = "spearman", use = "pairwise.complete.obs"), digits = 2),
corr = round(cor(usgs, usda, method = "pearson", use = "pairwise.complete.obs"), digits = 2),
n = n()) %>%
filter(n > 10) %>%
ungroup() %>%
arrange(methods, usgs_cmpd)By crop
stats_crop <- graph_data %>%
drop_na() %>%
group_by(CROP, usda_category, method) %>%
summarise(med_RE = round(median(RE, na.rm=T), digits = 1),
iqr_25 = round(quantile(RE, .25, na.rm=T), digits = 1),
iqr_75 = round(quantile(RE, .75, na.rm=T), digits = 1),
rank_corr = round(cor(usgs, usda, method = "spearman", use = "pairwise.complete.obs"), digits = 2),
corr = round(cor(usgs, usda, method = "pearson", use = "pairwise.complete.obs"), digits = 2),
n = n())By year
stats_year <- graph_data %>%
drop_na() %>%
group_by(YEAR, usda_category, method) %>%
summarise(med_RE = round(median(RE, na.rm=T), digits = 1),
iqr_25 = round(quantile(RE, .25, na.rm=T), digits = 1),
iqr_75 = round(quantile(RE, .75, na.rm=T), digits = 1),
rank_corr = round(cor(usgs, usda, method = "spearman", use = "pairwise.complete.obs"), digits = 2),
corr = round(cor(usgs, usda, method = "pearson", use = "pairwise.complete.obs"), digits = 2),
n = n())This R Markdown site was created with workflowr