Process USDA pesticide application data
Maggie Douglas
Last updated: 2021-12-09
Purpose
Process, clean, and organize data from USDA Chemical Use Survey so that it can be compared to estimates derived from USGS data.
Libraries & functions
library(tidyverse)
library(data.table)
library(vcd)
# function to fix USDA acreage data stored as character
acreNum<-function(x){
as.numeric(as.character(gsub(",", "", x)))
}Load data
qs.environment.pest.st <- fread("../output_big/nass_survey/qs.environment.pest.st_20200528.csv")
qs.environment.pest.st$VALUE <- acreNum(qs.environment.pest.st$VALUE)
str(qs.environment.pest.st)Classes 'data.table' and 'data.frame': 409142 obs. of 39 variables:
$ SOURCE_DESC : chr "SURVEY" "SURVEY" "SURVEY" "SURVEY" ...
$ SECTOR_DESC : chr "ENVIRONMENTAL" "ENVIRONMENTAL" "ENVIRONMENTAL" "ENVIRONMENTAL" ...
$ GROUP_DESC : chr "VEGETABLES" "VEGETABLES" "VEGETABLES" "VEGETABLES" ...
$ COMMODITY_DESC : chr "BEANS" "BEANS" "CARROTS" "SWEET CORN" ...
$ CLASS_DESC : chr "SNAP" "SNAP" "ALL CLASSES" "ALL CLASSES" ...
$ PRODN_PRACTICE_DESC : chr "ALL PRODUCTION PRACTICES" "ALL PRODUCTION PRACTICES" "ALL PRODUCTION PRACTICES" "ALL PRODUCTION PRACTICES" ...
$ UTIL_PRACTICE_DESC : chr "ALL UTILIZATION PRACTICES" "ALL UTILIZATION PRACTICES" "ALL UTILIZATION PRACTICES" "ALL UTILIZATION PRACTICES" ...
$ STATISTICCAT_DESC : chr "APPLICATIONS" "APPLICATIONS" "APPLICATIONS" "TREATED" ...
$ UNIT_DESC : chr "LB" "LB" "LB" "PCT OF AREA PLANTED, AVG" ...
$ SHORT_DESC : chr "BEANS, SNAP - APPLICATIONS, MEASURED IN LB" "BEANS, SNAP - APPLICATIONS, MEASURED IN LB" "CARROTS - APPLICATIONS, MEASURED IN LB" "SWEET CORN - TREATED, MEASURED IN PCT OF AREA PLANTED, AVG" ...
$ DOMAIN_DESC : chr "CHEMICAL, HERBICIDE" "CHEMICAL, HERBICIDE" "CHEMICAL, HERBICIDE" "CHEMICAL, HERBICIDE" ...
$ DOMAINCAT_DESC : chr "CHEMICAL, HERBICIDE: (TOTAL)" "CHEMICAL, HERBICIDE: (TOTAL)" "CHEMICAL, HERBICIDE: (TOTAL)" "CHEMICAL, HERBICIDE: (TOTAL)" ...
$ AGG_LEVEL_DESC : chr "STATE" "STATE" "STATE" "STATE" ...
$ STATE_ANSI : int 26 55 53 12 53 13 6 13 18 18 ...
$ STATE_FIPS_CODE : int 26 55 53 12 53 13 6 13 18 18 ...
$ STATE_ALPHA : chr "MI" "WI" "WA" "FL" ...
$ STATE_NAME : chr "MICHIGAN" "WISCONSIN" "WASHINGTON" "FLORIDA" ...
$ ASD_CODE : logi NA NA NA NA NA NA ...
$ ASD_DESC : logi NA NA NA NA NA NA ...
$ COUNTY_ANSI : logi NA NA NA NA NA NA ...
$ COUNTY_CODE : logi NA NA NA NA NA NA ...
$ COUNTY_NAME : logi NA NA NA NA NA NA ...
$ REGION_DESC : logi NA NA NA NA NA NA ...
$ ZIP_5 : logi NA NA NA NA NA NA ...
$ WATERSHED_CODE : int 0 0 0 0 0 0 0 0 0 0 ...
$ WATERSHED_DESC : logi NA NA NA NA NA NA ...
$ CONGR_DISTRICT_CODE : logi NA NA NA NA NA NA ...
$ COUNTRY_CODE : int 9000 9000 9000 9000 9000 9000 9000 9000 9000 9000 ...
$ COUNTRY_NAME : chr "UNITED STATES" "UNITED STATES" "UNITED STATES" "UNITED STATES" ...
$ LOCATION_DESC : chr "MICHIGAN" "WISCONSIN" "WASHINGTON" "FLORIDA" ...
$ YEAR : int 2018 2018 2018 2018 2018 2018 2018 2018 2018 2018 ...
$ FREQ_DESC : chr "ANNUAL" "ANNUAL" "ANNUAL" "ANNUAL" ...
$ BEGIN_CODE : int 0 0 0 0 0 0 0 0 0 0 ...
$ END_CODE : int 0 0 0 0 0 0 0 0 0 0 ...
$ REFERENCE_PERIOD_DESC: chr "YEAR" "YEAR" "YEAR" "YEAR" ...
$ WEEK_ENDING : logi NA NA NA NA NA NA ...
$ LOAD_TIME : POSIXct, format: "2018-07-24 15:00:08" "2018-07-24 15:00:08" ...
$ VALUE : num 68200 142600 NA NA 85 ...
$ CV_% : chr "" "" "" "" ...
- attr(*, ".internal.selfref")=<externalptr> crop_key <- read.csv("../keys/crop_key_summary.csv")
crop_key_CA <- read.csv("../keys/crop_key_summary_CA.csv")Reorganize application rate data
# select data on lb/year/state
lb <- qs.environment.pest.st %>%
filter(UNIT_DESC=="LB") %>%
select(STATE_ALPHA, STATE_ANSI, YEAR, COMMODITY_DESC, CLASS_DESC,
PRODN_PRACTICE_DESC, UTIL_PRACTICE_DESC,
DOMAIN_DESC, DOMAINCAT_DESC, UNIT_DESC, VALUE, 'CV_%') %>%
rename(lb = VALUE) %>%
select(-UNIT_DESC) %>%
na.omit()
# select data on lb/acre/application
app_rate <- qs.environment.pest.st %>%
filter(UNIT_DESC=="LB / ACRE / APPLICATION, AVG") %>%
select(STATE_ALPHA, STATE_ANSI, YEAR, COMMODITY_DESC, CLASS_DESC,
PRODN_PRACTICE_DESC, UTIL_PRACTICE_DESC,
DOMAIN_DESC, DOMAINCAT_DESC, UNIT_DESC, VALUE, 'CV_%') %>%
rename(lb_ac_yr_app_ave = VALUE) %>%
select(-UNIT_DESC) %>%
na.omit()
# select data on lb/acre/year total
app_rate_tot <- qs.environment.pest.st %>%
filter(UNIT_DESC=="LB / ACRE / YEAR, AVG") %>%
select(STATE_ALPHA, STATE_ANSI, YEAR, COMMODITY_DESC, CLASS_DESC,
PRODN_PRACTICE_DESC, UTIL_PRACTICE_DESC,
DOMAIN_DESC, DOMAINCAT_DESC, UNIT_DESC, VALUE, 'CV_%') %>%
rename(lb_ac_yr_ave = VALUE) %>%
select(-UNIT_DESC) %>%
na.omit()
# select data on percent treated
# focused on bearing acreage b/c only one observation for non-bearing!
perc_trt <- qs.environment.pest.st %>%
filter(UNIT_DESC %in% c("PCT OF AREA PLANTED, AVG",
"PCT OF AREA BEARING, AVG")) %>%
select(STATE_ALPHA, STATE_ANSI, YEAR, COMMODITY_DESC, CLASS_DESC,
PRODN_PRACTICE_DESC, UTIL_PRACTICE_DESC,
DOMAIN_DESC, DOMAINCAT_DESC, UNIT_DESC, VALUE, 'CV_%') %>%
rename(perc_trt_ave = VALUE) %>%
select(-UNIT_DESC) %>%
na.omit()
# combine datasets and calculate application rates for individual compounds
comb_data <- app_rate_tot %>%
full_join(perc_trt, by = c("STATE_ALPHA", "STATE_ANSI", "YEAR", "COMMODITY_DESC",
"CLASS_DESC", "PRODN_PRACTICE_DESC", "UTIL_PRACTICE_DESC",
"DOMAIN_DESC", "DOMAINCAT_DESC")) %>%
full_join(lb, by = c("STATE_ALPHA", "STATE_ANSI", "YEAR", "COMMODITY_DESC",
"CLASS_DESC", "PRODN_PRACTICE_DESC", "UTIL_PRACTICE_DESC",
"DOMAIN_DESC", "DOMAINCAT_DESC")) %>%
filter(!grepl("(TOTAL)", DOMAINCAT_DESC), # filter out aggregate values, post harvest, and NAs for main indicator
UTIL_PRACTICE_DESC != "POST HARVEST",
PRODN_PRACTICE_DESC != "ORGANIC",
!is.na(lb_ac_yr_ave)) %>%
# drop_na %>%
mutate(lb_ac_yr_ave = lb_ac_yr_ave*(perc_trt_ave/100), # calculate annual ave application on all acres
kg_ha_yr_ave = lb_ac_yr_ave*2.47105*.453592) # convert to kg/ha
# generate dataset for aggregate indicators
agg_data <- app_rate_tot %>%
full_join(perc_trt, by = c("STATE_ALPHA", "STATE_ANSI", "YEAR", "COMMODITY_DESC",
"CLASS_DESC", "PRODN_PRACTICE_DESC", "UTIL_PRACTICE_DESC",
"DOMAIN_DESC", "DOMAINCAT_DESC")) %>%
full_join(lb, by = c("STATE_ALPHA", "STATE_ANSI", "YEAR", "COMMODITY_DESC",
"CLASS_DESC", "PRODN_PRACTICE_DESC", "UTIL_PRACTICE_DESC",
"DOMAIN_DESC", "DOMAINCAT_DESC")) %>%
filter(grepl("(TOTAL)", DOMAINCAT_DESC), # filter out unwanted values
UTIL_PRACTICE_DESC != "POST HARVEST",
PRODN_PRACTICE_DESC != "ORGANIC") %>%
mutate(kg = lb*0.453592) %>%
select(STATE_ALPHA:DOMAINCAT_DESC, perc_trt_ave, lb, kg)Check some patterns - e.g. glyphosate
test <- comb_data %>%
filter(COMMODITY_DESC == "SOYBEANS",
grepl("GLYPHOSATE", DOMAINCAT_DESC)) %>%
group_by(STATE_ALPHA, YEAR, COMMODITY_DESC, CLASS_DESC,
PRODN_PRACTICE_DESC, UTIL_PRACTICE_DESC,
DOMAIN_DESC) %>%
summarise(kg_ha_yr_sum = sum(kg_ha_yr_ave))
ggplot(data = test, aes(group = YEAR, x = YEAR, y = kg_ha_yr_sum)) +
geom_boxplot()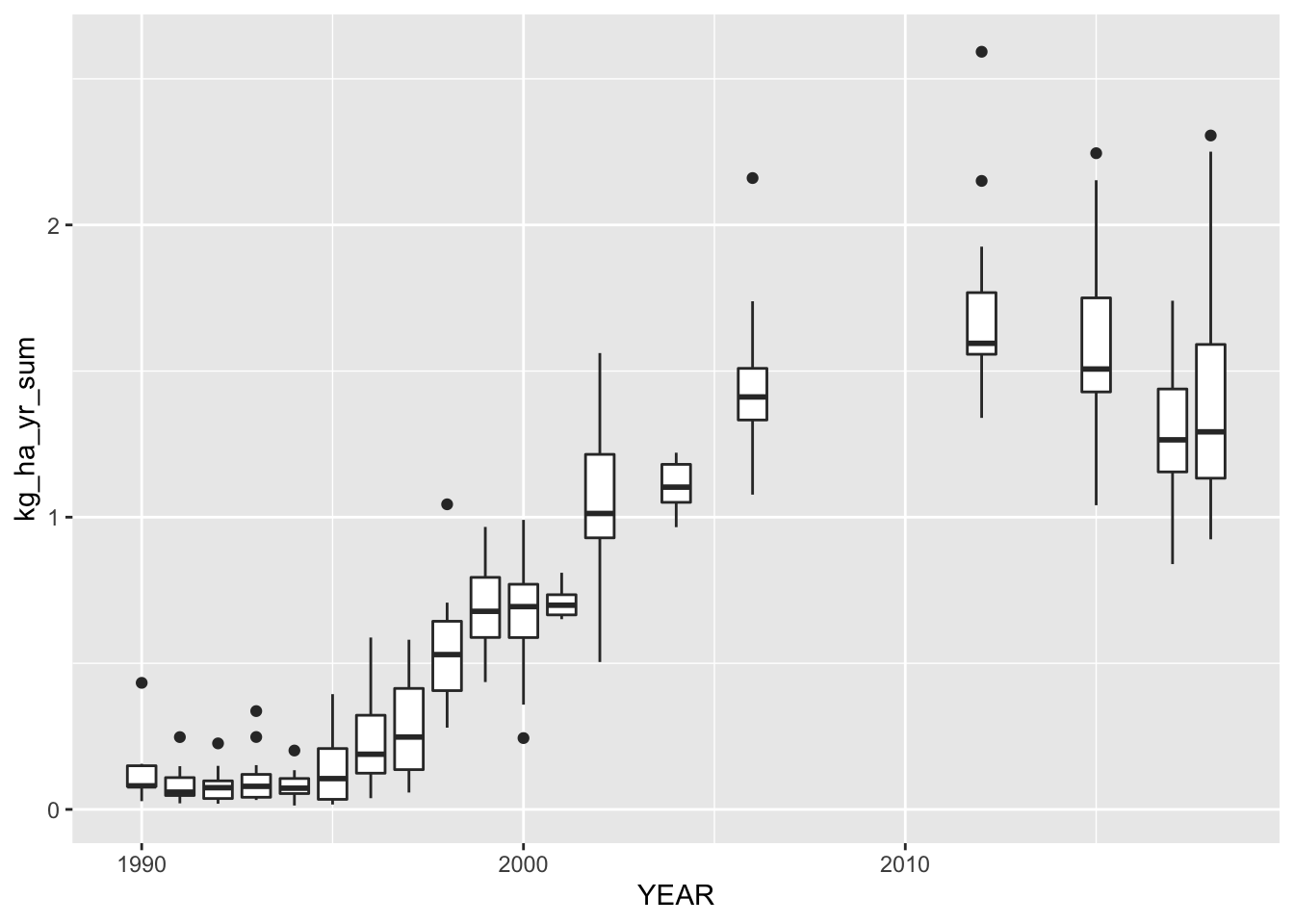
ggplot(data = test, aes(x = YEAR, y = kg_ha_yr_sum)) +
geom_line() +
facet_wrap(~STATE_ALPHA)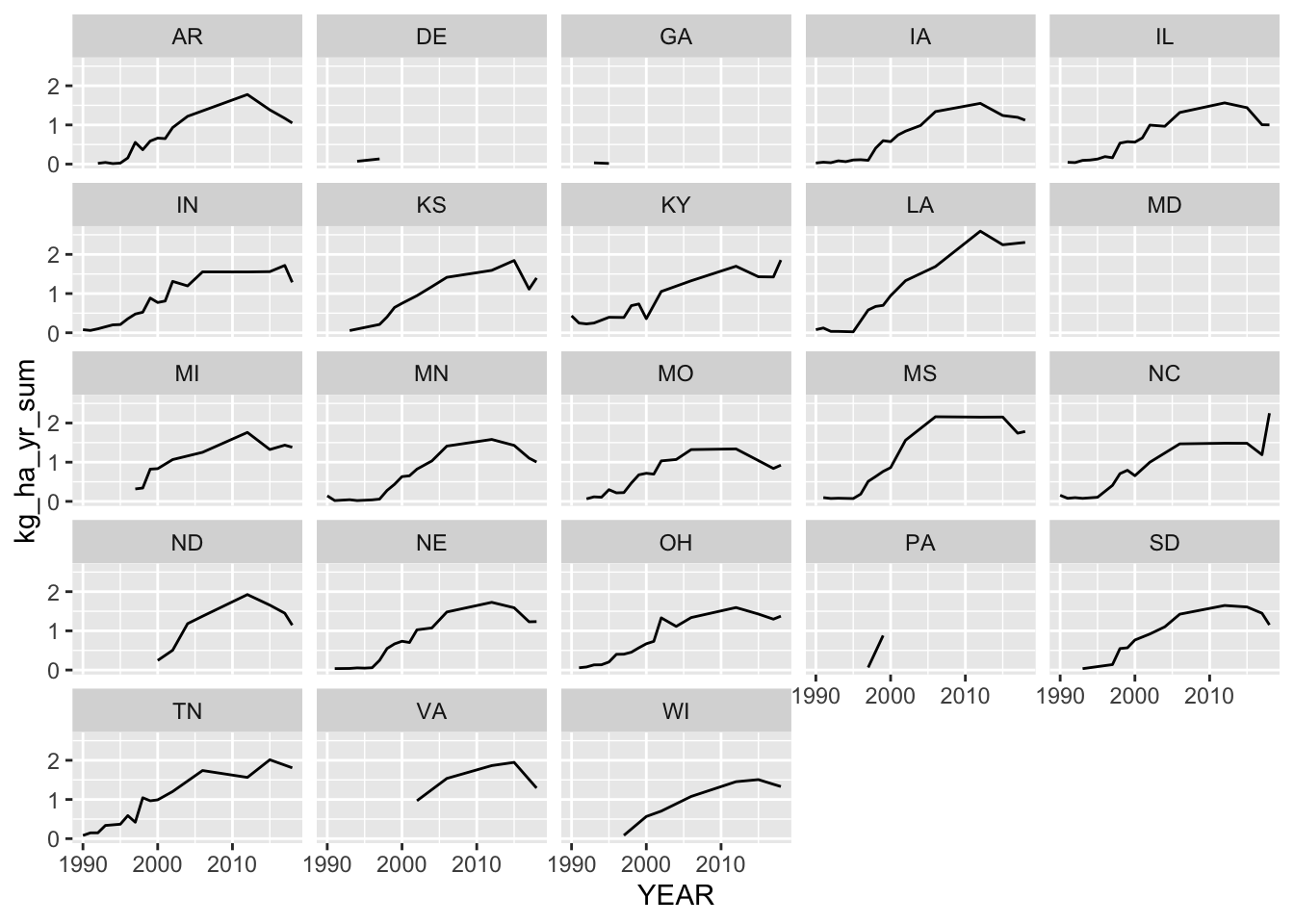
Check distribution of observations across crops
# number of state-year observations for combos of commodity + AI
data_dist_crop <- comb_data %>%
group_by(COMMODITY_DESC, DOMAIN_DESC) %>%
summarise(n = n()) %>%
arrange(-n)
agg_dist_crop <- agg_data %>%
group_by(COMMODITY_DESC, DOMAIN_DESC) %>%
summarise(n = n()) %>%
arrange(-n)
# Application rate data
ggplot(data = data_dist_crop, aes(x = COMMODITY_DESC, y = n)) +
geom_bar(stat = "identity") +
facet_grid(DOMAIN_DESC~.) +
theme_bw() +
theme(axis.text.x = element_text(angle = 90),
axis.title.x = element_blank()) 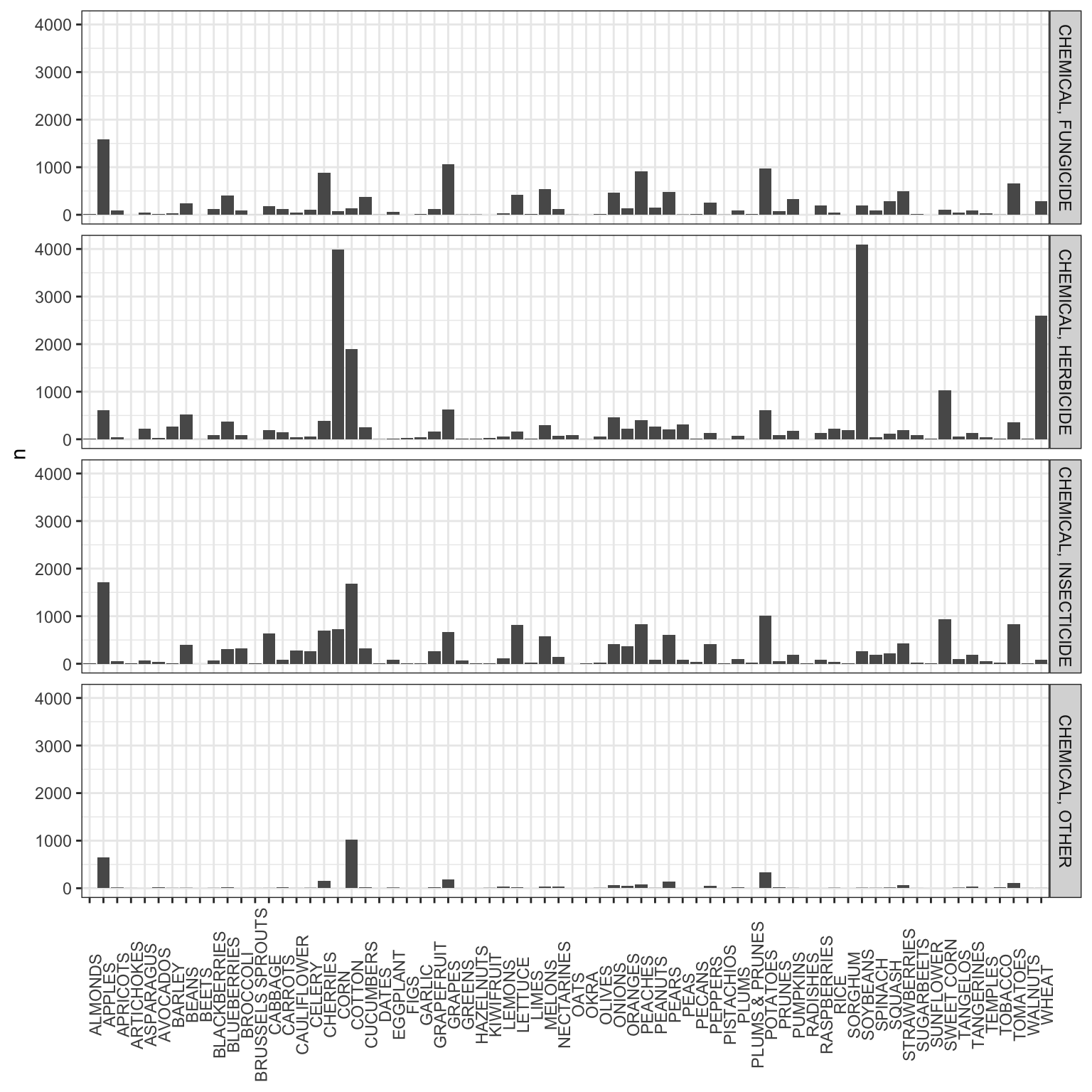
# Percent treated and total pounds applied data
ggplot(data = agg_dist_crop, aes(x = COMMODITY_DESC, y = n)) +
geom_bar(stat = "identity") +
facet_grid(DOMAIN_DESC~.) +
theme_bw() +
theme(axis.text.x = element_text(angle = 90),
axis.title.x = element_blank()) 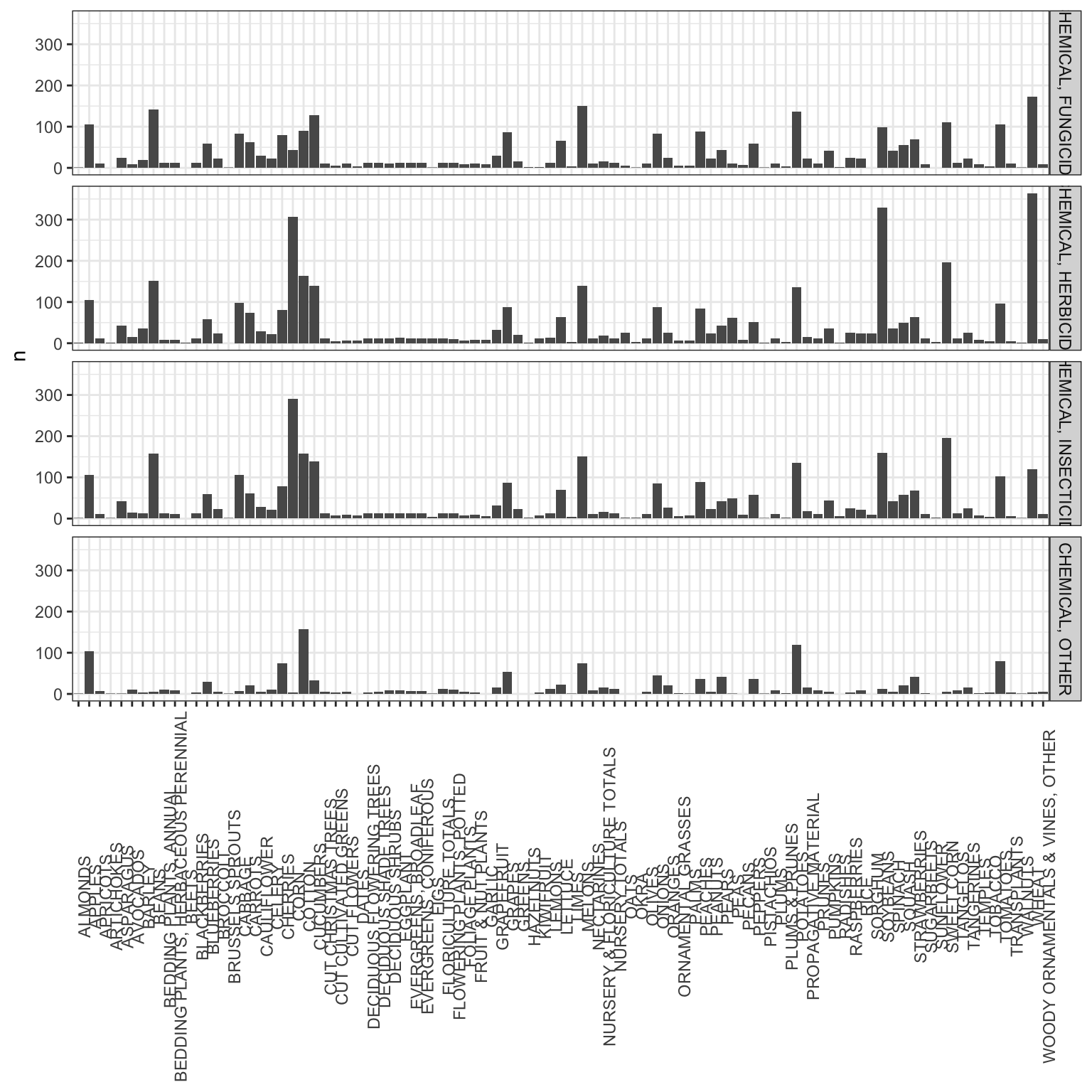
# examine only crops that closely match USGS categories
data_dist_main <- comb_data %>%
filter(COMMODITY_DESC %in% c("COTTON", "CORN", "SOYBEANS", "RICE", "WHEAT")) %>%
group_by(COMMODITY_DESC, CLASS_DESC, PRODN_PRACTICE_DESC,
UTIL_PRACTICE_DESC, DOMAIN_DESC, DOMAINCAT_DESC) %>%
summarise(n = n()) %>%
arrange(-n)
agg_dist_main <- agg_data %>%
filter(COMMODITY_DESC %in% c("COTTON", "CORN", "SOYBEANS", "RICE", "WHEAT")) %>%
group_by(COMMODITY_DESC, CLASS_DESC, PRODN_PRACTICE_DESC,
UTIL_PRACTICE_DESC, DOMAIN_DESC, DOMAINCAT_DESC) %>%
summarise(n = n()) %>%
arrange(-n)
# check for overlapping observations
dupes <- select(agg_data, YEAR, STATE_ALPHA, COMMODITY_DESC, CLASS_DESC, PRODN_PRACTICE_DESC, UTIL_PRACTICE_DESC) %>%
unique() %>%
group_by(YEAR, STATE_ALPHA, COMMODITY_DESC) %>%
summarise(n = n()) %>%
filter(n > 1) %>%
mutate(st_yr_cr = paste(YEAR, STATE_ALPHA, COMMODITY_DESC)) %>%
arrange(-n)
crops_dupe <- unique(dupes$st_yr_cr)
data_items <- select(agg_data, YEAR, STATE_ALPHA, COMMODITY_DESC,
CLASS_DESC, PRODN_PRACTICE_DESC, UTIL_PRACTICE_DESC) %>%
mutate(st_yr_cr = paste(YEAR, STATE_ALPHA, COMMODITY_DESC)) %>%
filter(YEAR > 1996 & YEAR < 2018 & st_yr_cr %in% crops_dupe) %>%
unique() %>%
arrange(COMMODITY_DESC, STATE_ALPHA, YEAR)
table(data_dist_main$PRODN_PRACTICE_DESC)
ALL PRODUCTION PRACTICES
618 table(data_dist_main$UTIL_PRACTICE_DESC)
ALL UTILIZATION PRACTICES
618 table(data_dist_main$CLASS_DESC)
ALL CLASSES SPRING, (EXCL DURUM) SPRING, DURUM
327 60 42
UPLAND WINTER
119 70 Filter dataset to those in USGS dataset
# focus on crops with a species match in the USGS dataset
comb_data_usgs <- comb_data %>%
filter(COMMODITY_DESC %in% c("COTTON", "CORN", "SOYBEANS", "RICE", "WHEAT"),
PRODN_PRACTICE_DESC == "ALL PRODUCTION PRACTICES",
UTIL_PRACTICE_DESC == "ALL UTILIZATION PRACTICES") %>%
select(-PRODN_PRACTICE_DESC, -UTIL_PRACTICE_DESC)
comb_data_usgs_class <- comb_data_usgs %>%
group_by(STATE_ALPHA, COMMODITY_DESC, CLASS_DESC) %>%
summarise(n = n())
comb_data_class <- comb_data %>%
group_by(STATE_ALPHA, COMMODITY_DESC, CLASS_DESC) %>%
summarise(n = n())
table(comb_data_usgs$COMMODITY_DESC,comb_data_usgs$CLASS_DESC)
ALL CLASSES SPRING, (EXCL DURUM) SPRING, DURUM UPLAND WINTER
CORN 4775 0 0 0 0
COTTON 1212 0 0 3522 0
RICE 314 0 0 0 0
SOYBEANS 4550 0 0 0 0
WHEAT 0 932 296 0 1744# same operation for aggregate data
agg_data_usgs <- agg_data %>%
filter(COMMODITY_DESC %in% c("COTTON", "CORN", "SOYBEANS", "RICE", "WHEAT"),
PRODN_PRACTICE_DESC == "ALL PRODUCTION PRACTICES",
UTIL_PRACTICE_DESC == "ALL UTILIZATION PRACTICES") %>%
select(-PRODN_PRACTICE_DESC, -UTIL_PRACTICE_DESC)
agg_data_class <- agg_data_usgs %>%
group_by(STATE_ALPHA, COMMODITY_DESC, CLASS_DESC) %>%
summarise(n = n())
table(agg_data_usgs$COMMODITY_DESC,agg_data_usgs$CLASS_DESC)
ALL CLASSES SPRING, (EXCL DURUM) SPRING, DURUM UPLAND WINTER
CORN 644 0 0 0 0
COTTON 143 0 0 427 0
RICE 77 0 0 0 0
SOYBEANS 600 0 0 0 0
WHEAT 0 160 43 0 456Save files
write.csv(comb_data, "../output_big/nass_survey/qs.environment.pest.st.app.rate_20200528.csv", row.names=F)
write.csv(comb_data_usgs, "../output_big/nass_survey/qs.environment.pest.st.app.rate.usgs_20200528.csv", row.names=F)
write.csv(agg_data, "../output_big/nass_survey/qs.environment.pest.st.agg_20200528.csv", row.names=F)
write.csv(agg_data_usgs, "../output_big/nass_survey/qs.environment.pest.st.agg.usgs_20200528.csv", row.names=F)Session information
sessionInfo()R version 3.6.1 (2019-07-05)
Platform: x86_64-apple-darwin15.6.0 (64-bit)
Running under: macOS Catalina 10.15.7
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/3.6/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/3.6/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] grid stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] vcd_1.4-4 data.table_1.14.0 forcats_0.4.0
[4] stringr_1.4.0 dplyr_0.8.3 purrr_0.3.2
[7] readr_1.3.1 tidyr_1.1.0 tibble_2.1.3
[10] ggplot2_3.2.0 tidyverse_1.2.1
loaded via a namespace (and not attached):
[1] zoo_1.8-6 tidyselect_1.1.0 xfun_0.8 reshape2_1.4.3
[5] haven_2.1.1 lattice_0.20-38 colorspace_1.4-1 vctrs_0.3.2
[9] generics_0.0.2 htmltools_0.3.6 yaml_2.2.0 rlang_0.4.7
[13] pillar_1.4.2 glue_1.3.1 withr_2.1.2 modelr_0.1.4
[17] readxl_1.3.1 plyr_1.8.4 lifecycle_0.2.0 munsell_0.5.0
[21] gtable_0.3.0 cellranger_1.1.0 rvest_0.3.4 evaluate_0.14
[25] labeling_0.3 knitr_1.23 lmtest_0.9-37 broom_0.5.2
[29] Rcpp_1.0.6 scales_1.0.0 backports_1.1.4 jsonlite_1.6
[33] hms_0.5.0 digest_0.6.20 stringi_1.4.3 cli_1.1.0
[37] tools_3.6.1 magrittr_1.5 lazyeval_0.2.2 crayon_1.3.4
[41] pkgconfig_2.0.2 MASS_7.3-51.4 xml2_1.2.0 lubridate_1.7.4
[45] assertthat_0.2.1 rmarkdown_1.14 httr_1.4.2 rstudioapi_0.10
[49] R6_2.4.0 nlme_3.1-140 compiler_3.6.1 This R Markdown site was created with workflowr